Essential Peptide Optimization Strategies for Therapeutic Success
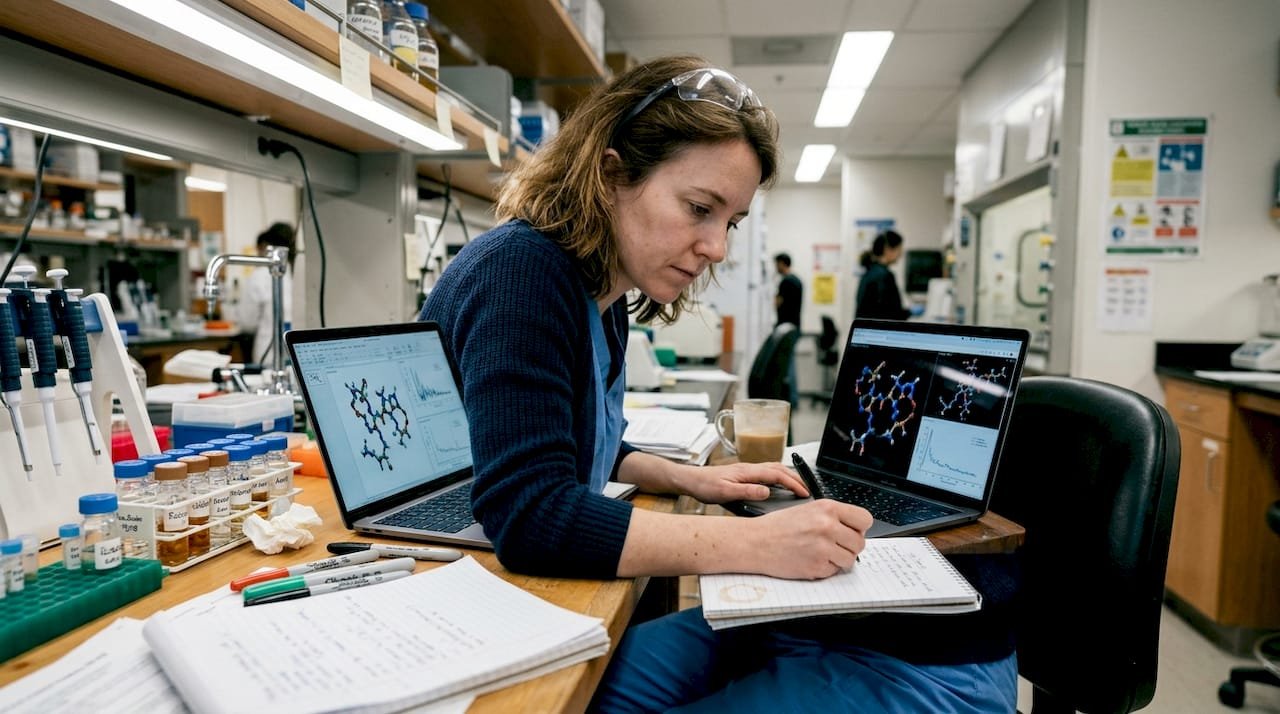

TL;DR:
- Peptide optimization involves balancing stability, bioavailability, selectivity, and safety trade-offs.
- Combining chemical modifications with AI-driven design enhances stability and efficacy more effectively.
- Empirical validation remains essential despite advances in computational tools and in silico predictions.
Peptide optimization is never a single-variable problem. Even experienced R&D teams wrestle with the competing demands of stability, bioavailability, selectivity, and translational relevance simultaneously. Push solubility too far and you risk aggregation. Maximize protease resistance and you may trigger immunogenicity. The field has matured considerably, and the best teams now draw on chemical, computational, and hybrid methods in deliberate combination. This guide breaks down the core strategies, compares their strengths and limitations, and offers a decision-oriented framework grounded in current data.
Table of Contents
- Core criteria in peptide optimization
- Chemical modification methods: from substitution to lipidation
- Computational design and AI in peptide optimization
- Hybrid and integrated optimization: bridging chemistry and computation
- Edge cases, limitations, and best practices
- Why integration, not just innovation, drives peptide therapeutic progress
- Take your peptide optimization further with PrimeGen Labs
- Frequently asked questions
Key Takeaways
| Point | Details |
|---|---|
| Balance trade-offs | Effective peptide optimization requires balancing stability, efficacy, and delivery challenges. |
| Leverage hybrid methods | Integrating chemical modifications with computational design produces superior therapeutic candidates. |
| Validate experimentally | Even advanced AI-driven designs must be validated in biological systems for real-world effectiveness. |
| Anticipate limitations | Understanding risks like aggregation and immunogenicity helps avoid costly peptide development failures. |
Core criteria in peptide optimization
With a clear understanding of why advanced optimization matters, let’s break down the criteria that drive strategy selection.
Every optimization decision begins with defining the target profile: the specific combination of stability, bioavailability, target specificity, and pharmacokinetic or pharmacodynamic (PK/PD) behavior that the therapeutic candidate must achieve. These properties are deeply interconnected, and improving one often compromises another.
Consider a few foundational trade-offs that every peptide researcher encounters:
- Stability vs. immunogenicity. Introducing non-natural amino acids or lipid chains dramatically extends half-life, but foreign structural elements raise immunogenic risk.
- Hydrophobicity vs. aggregation. Lipidation and certain substitutions increase plasma binding, yet excess hydrophobicity causes self-association and formulation failure.
- Potency vs. specificity. Optimization for receptor affinity can erode selectivity across closely related receptor subtypes, complicating safety profiling.
- In vitro performance vs. in vivo translation. Plasma stability assays are predictive but not perfectly translatable, especially when species-specific metabolic factors differ.
“Unmodified peptides may degrade in minutes; stabilized analogs can exceed 100-fold increases in half-life.”
Unmodified peptides show a half-life of roughly 2 to 30 minutes in plasma, while semaglutide achieves approximately 165 hours via lipidation and cyclic peptides routinely demonstrate more than 100-fold stability increases in plasma assays. These benchmarks are essential reference points when evaluating whether a modification strategy is achieving meaningful therapeutic progress.
Pro Tip: Computational predictions are valuable starting points, but they cannot replace empirical plasma stability assays and in vivo PK studies. Always build validation checkpoints into your optimization workflow before advancing a candidate.
Also consider reviewing safety tips for peptide optimization to ensure your R&D process accounts for both performance and risk.
Chemical modification methods: from substitution to lipidation
Having established what to evaluate, the next step is understanding which chemical modifications best serve those goals.
Chemical modification strategies including amino acid substitution, lipidation, PEGylation, cyclization, and glycosylation are the primary toolkit for enhancing stability, improving PK properties, and boosting therapeutic efficacy by regulating charge, hydrophobicity, conformation, and sequence integrity.
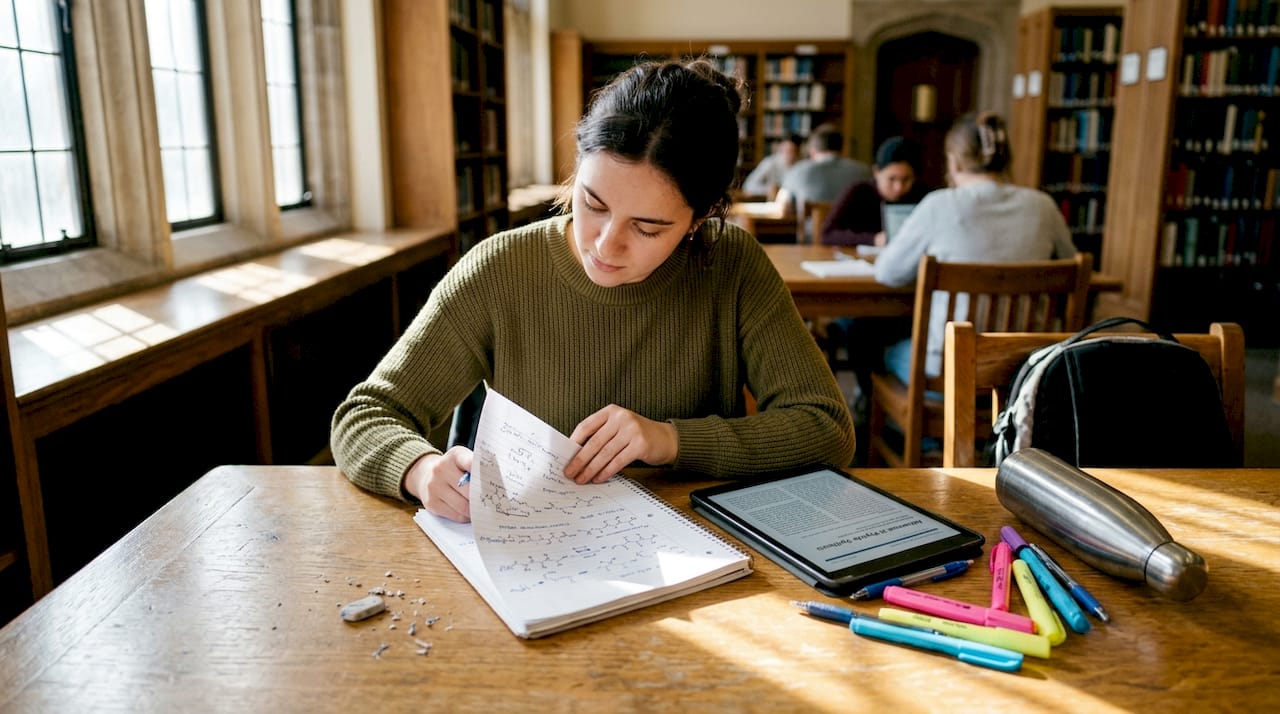

Here is how the major approaches compare across key performance properties:
| Modification | Half-life gain | Protease resistance | Solubility impact | Key example |
|---|---|---|---|---|
| Lipidation | Very high | Moderate | Variable | Semaglutide (~165h) |
| PEGylation | High | Moderate | Increased | PEGylated insulin analogs |
| Cyclization | High | Very high | Reduced | Cyclosporine A |
| D-amino acid substitution | Moderate | High | Neutral | Multiple AMP analogs |
| Glycosylation | Moderate | Low | Increased | EPO glycopeptides |
Some quantitative highlights:
- Semaglutide achieves a half-life of ~165 hours versus liraglutide’s ~13 hours, driven primarily by its C18 fatty diacid chain enabling tighter albumin binding.
- Cyclic peptides in plasma stability assays regularly exceed 100-fold half-life increases compared to linear counterparts.
- Lipidation promotes albumin binding, extending half-life from minutes to days or weeks, while PEGylation increases hydrodynamic radius, directly reducing renal clearance.
For researchers working on oral or subcutaneous delivery, understanding the downstream effects on peptide bioavailability strategies is a critical parallel workstream.
Pro Tip: Hybrid modification approaches, such as combining lipidation with cyclization, can address stability-solubility trade-offs that neither method resolves alone. Combining improving peptide outcomes approaches is more effective than relying on a single modification.
Computational design and AI in peptide optimization
As chemical modifications mature, innovators are increasingly leveraging computational power, especially AI, to push optimization further.
Modern computational strategies have moved well beyond basic homology modeling. The core techniques now in active use include:
- Molecular docking. Generates binding pose predictions between peptide candidates and target proteins, enabling rapid screening of sequence variants.
- Molecular dynamics (MD) simulations. Assesses structural flexibility and stability over time, critical for flagging aggregation-prone regions.
- Free energy calculations. Quantifies binding affinity differences between analogs, supporting lead selection before synthesis.
- AI-driven generative design. Uses machine learning models to propose novel sequences optimized for target properties.
Computational strategies including AI-driven tools like AlphaFold, Rosetta, EvoPepFold, PepMimic, and KCM enable rational peptide design, binding pose generation, flexibility assessment, and affinity optimization at a scale no chemistry team can match manually.
| Tool | Primary input | Key output | Best application |
|---|---|---|---|
| AlphaFold | Sequence | 3D structure prediction | Structural modeling |
| Rosetta | Structure/sequence | Energy-minimized designs | Stability optimization |
| EvoPepFold | GA + AlphaFold/Rosetta | Improved interaction energy | Binding affinity refinement |
| PepMimic | Target binding site | Peptidomimetics | PD-L1, cancer targets |
| KCM | Sequence library | Antimicrobial candidates | Infectious disease |
AI-guided methods like EvoPepFold improve interaction energy scores systematically, while PepMimic achieves binding affinities in the KD 10^-9 M range for targets such as PD-L1, and KCM designs antimicrobial peptides with demonstrated in vivo efficacy.
The practical limitation worth flagging: AI tools struggle with high-flexibility peptides where the conformational ensemble is broad and poorly captured by static docking. Experimental validation remains non-negotiable regardless of in silico confidence scores. Stay current with peptide research trends to track how these tools evolve alongside wet-lab methods.
Hybrid and integrated optimization: bridging chemistry and computation
Instead of treating chemistry and computation as separate silos, leading groups now synthesize both approaches in one workflow.
Hybrid approaches combining chemical modifications with computational design consistently yield superior therapeutic candidates. AI accelerates the exploration of sequence space, but it requires experimental validation to account for the real-world biological complexity that docking algorithms cannot fully model.
A practical integrated workflow looks like this:
- Step 1: AI-driven pre-screening. Use tools like EvoPepFold or Rosetta to narrow a candidate library from thousands to dozens of high-affinity sequences.
- Step 2: Chemical modification layering. Apply targeted modifications, for example cyclization to address protease vulnerability or lipidation to extend half-life, based on the structural insights from Step 1.
- Step 3: Plasma stability assays. Run in vitro validation to confirm that predicted gains translate to measurable half-life improvements.
- Step 4: PK/PD profiling. Progress candidates with validated stability and selectivity into animal PK studies before investing in full synthesis scale-up.
- Step 5: Iterative refinement. Feed empirical data back into the computational model to refine next-generation designs.
A strong illustration: pairing EvoPepFold-guided sequence optimization with downstream cyclization consistently improves protease resistance beyond what either approach achieves independently. For details on safe laboratory practices during scale-up, peptide reconstitution safety is essential reading before moving into formulation work.
Pro Tip: Multi-modification strategies that combine AI-selected sequences with chemical engineering address stability-solubility trade-offs far more effectively than standalone single-method approaches.
Edge cases, limitations, and best practices
No plan is complete unless it accounts for real-world exceptions and limitations that upend even the best theoretical designs.
Common edge cases in peptide optimization include high-flexibility peptides with poor oral bioavailability, aggregation driven by excess hydrophobicity following lipidation, immunogenicity risks from unnatural modifications, and significant species differences in metabolic stability between rat and human systems.
Here is a numbered framework for anticipating and managing them:
- High-flexibility peptides. These resist accurate docking predictions. Mitigation: Use MD simulations to map the conformational ensemble before selecting modifications. Cyclization is often the most practical structural fix.
- Aggregation risk. Excess hydrophobicity from lipid chains or beta-sheet-prone sequences drives self-association. Mitigation: Monitor critical aggregation concentration (CAC) during formulation screening. PEGylation or solubilizing residue insertion can break aggregation propensity.
- Immunogenicity. Unnatural amino acids and large PEG chains can generate anti-drug antibodies. Mitigation: Screen immunogenic epitopes computationally using tools like NetMHCpan early in the design cycle rather than waiting for in vivo studies.
- Species translation failures. Rat plasma proteases differ meaningfully from human, making rat stability data an incomplete predictor of human PK. Mitigation: Include human plasma stability assays in parallel with rodent studies from the outset.
- Off-target receptor activity. Modifications that increase receptor affinity can expand binding across related subtypes. Mitigation: Run selectivity panels early, not just after lead selection.
“Even the most promising in silico designs demand in vivo validation against species-specific metabolic factors.”
For additional guidance on managing these complexities in practice, safe peptide dosing protocols provide a structured reference.
Why integration, not just innovation, drives peptide therapeutic progress
The field has a recurring tendency to chase the next breakthrough in isolation. When AlphaFold emerged, many teams pivoted to purely computational workflows. When lipidation produced semaglutide’s remarkable half-life numbers, others pursued lipidation as a universal solution. Neither instinct is wrong in itself, but both miss the more durable point.
The highest-performing programs we observe share one consistent feature: they treat chemistry, computation, and empirical validation as a closed loop rather than sequential stages. Computational tools cut synthesis costs and accelerate candidate selection. Chemical modifications deliver properties that no algorithm alone can engineer into a sequence. And experimental validation catches the failures that even excellent models will inevitably generate.
The contrarian view worth holding: overselling AI-driven design without grounding it in chemical context wastes both time and resources. A KD of 10^-9 M in silico means nothing if the sequence aggregates in formulation. Evidence for peptide performance always has to be empirical at some stage. The teams that internalize this early move faster, not slower, because they avoid rebuilding after late-stage failures.
Take your peptide optimization further with PrimeGen Labs
If you’re ready to turn insights into action, here’s how we can support your next discovery.
PrimeGen Labs is built to support researchers who need more than off-the-shelf answers. Whether you’re working through a stability challenge, evaluating modification strategies, or scaling a candidate toward clinical relevance, our resources are designed to meet you at the right stage of your workflow.
Start with our comprehensive peptide optimization guide for a grounded overview of performance-oriented approaches, or explore the full peptide product catalog to find candidates aligned with your research targets. For a research-backed review of outcomes and risk profiles, our peptide evidence and benefits resource is the right starting point.
Frequently asked questions
What is the most effective chemical modification for extending peptide half-life?
Lipidation is the most effective modification for extending half-life, with semaglutide reaching approximately 165 hours via lipidation compared to the 2 to 30 minutes typical of unmodified peptides.
Which AI tool is best for peptide optimization?
EvoPepFold leads for binding affinity improvement by combining genetic algorithms with AlphaFold and Rosetta scoring, but all AI outputs require experimental validation before advancing candidates.
What are the main risks of peptide modification?
The primary risks are aggregation, immunogenicity, and species-specific stability differences, each of which requires targeted mitigation during the design and formulation phases.
Why combine chemical and computational strategies?
Hybrid approaches yield superior therapeutic candidates because AI accelerates candidate selection while chemical modifications deliver properties that computational models alone cannot engineer or guarantee in a biological environment.
